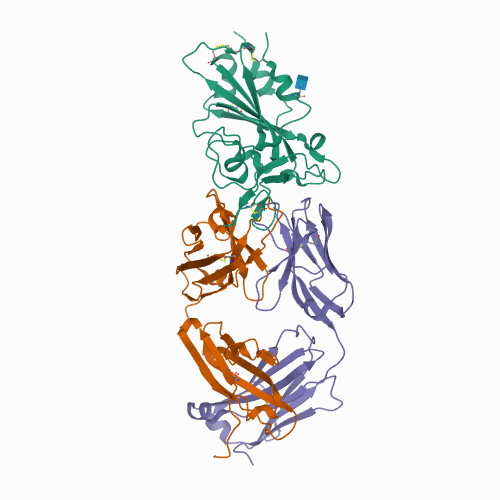
SARS-CoV-2 evolution in an immunocompromised host reveals shared neutralization escape mechanisms.
Publication Type:
Journal ArticleSource:
Cell, Volume 184, Issue 10 (2021)Abstract:
<p style="margin-top: 0px; margin-bottom: 0px; font-stretch: normal; font-size: 12px; line-height: normal; font-family: Helvetica;">Many individuals mount nearly identical antibody responses to SARS-CoV-2. To gain insight into how the viral spike (S) protein receptor-binding domain (RBD) might evolve in response to common antibody responses, we studied mutations occurring during virus evolution in a persistently infected immunocompromised individual. We use antibody Fab/RBD structures to predict, and pseudotypes to confirm, that mutations found in late-stage evolved S variants confer resistance to a common class of SARS-CoV-2 neutralizing antibodies we isolated from a healthy COVID-19 convalescent donor. Resistance extends to the polyclonal serum immunoglobulins of four out of four healthy convalescent donors we tested and to monoclonal antibodies in clinical use. We further show that affinity maturation is unimportant for wild-type virus neutralization but is critical to neutralization breadth. Because the mutations we studied foreshadowed emerging variants that are now circulating across the globe, our results have implications to the long-term efficacy of S-directed countermeasures.</p>